DIVISION POLYMER BIOMATERIALS SCIENCE
Bacterial Shape/Bacterials Mechanics
Independent Research associates & joint labs

The elementary unit of life is the cell. All cells are bounded by envelopes that confer size and shape, fundamental properties that are essential to many biological processes. One of the fundamental questions in cell biology is: If these shapes are required for proper physiology, how do the shapes themselves arise? My research (together with my collaborators) combines experiment and theory to identify the fundamental principles underlying the robust design of cell shape and the feedback mechanisms responsible for shape regulation to provide a comprehensive understanding of cellular growth and morphology across distant domains of life.
Soft Lithography
We are using soft lithography tools to design and fabricate microstructures and microfluidic chambers to study bacteria cell biology. With those tools at hand, we are manipulating cell shape, confine spheroplasts and use microfluidics to investigate spatio-temporal bacterial dynamics.
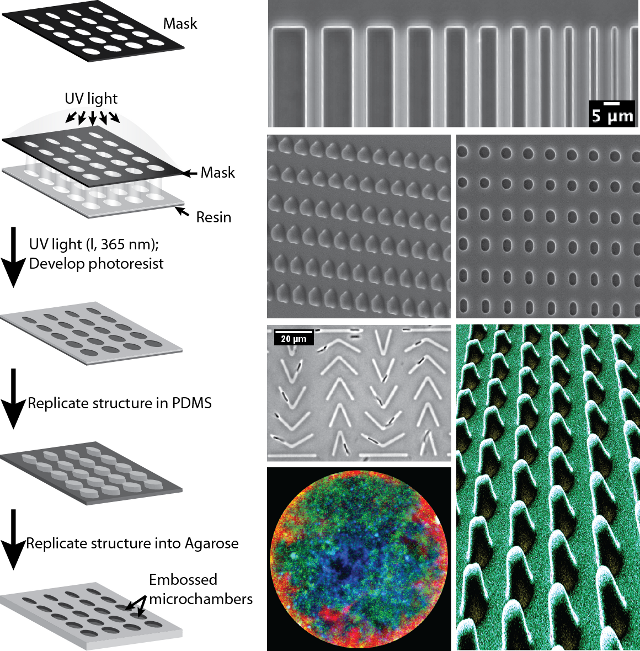
Bacterial cell shape regulation and mechanics
We designed toroidal microchambers to study shape recovery of rod-shaped bacteria after mechanical deformation. Bacterial cells are confined into the chambers and filamented before being released to analyse their recovery behaviour. We combine these mechanistics studies with protein localization at the single cell level using high-resolution microscopy.
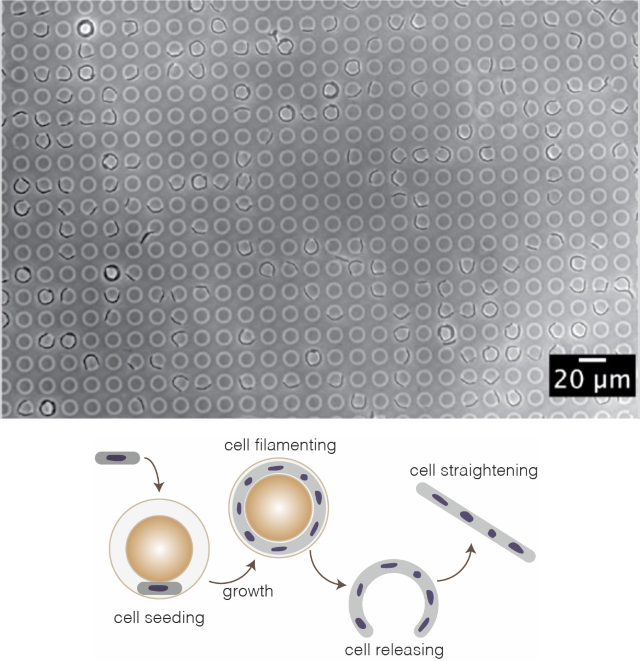

Bacterial cell size homeostasis
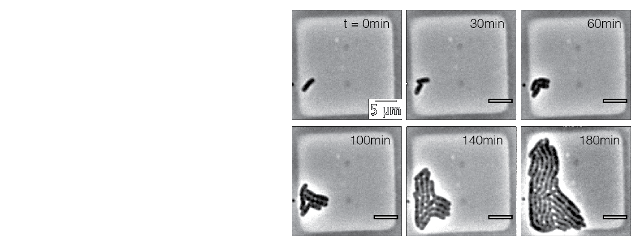
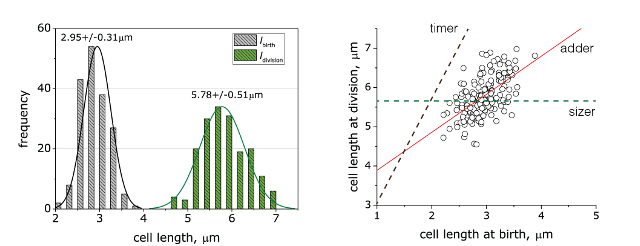
In collaboration with the Amir, Garner and Schmid lab, we are studying cell size homeostasis in E. coli using microchambers to follow growth at the single cell level. For example, a timelapse of growth for a single E. coli cell in microchambers over 5 generations is shown below. Histogram of E. coli cell lengths shows narrow distributions for birth (grey) and division (green), respectively. Cell lengths at birth and division are linearly related, and linear regression of the raw data (solid red line) suggests that the cells are growing following to the the adder mechanism.

Our collaborators
SELECTED REFERENCES (see google scholar)
Wong F., Wilson S., Helbig R., Hegde S., Aftenieva O., Zheng H., Liu C., Pilizota T., Garner E.C., Amir A., Renner L.D. Understanding Beta-Lactam-Induced Lysis at the Single-Cell Level. Frontiers in Microbiology 12:712007 (2021).
Wong F., Stokes J.M., Cervantes B., Penkov S., Friedrichs J., Renner L.D., Collins J.J. Cytoplasmic condensation induced by membrane damage is associated with antibiotic lethality. Nature Communications 12:1-15 (2021).
Hussain S., Wivagg C.N., Szwedziak P., Wong F., Schaefer K., Izoré T., Renner L.D., Holmes M.J., Sun Y., Bisson-Filho A.W., Walker S., Amir A., Löwe J., Garner E.C. MreB filaments align along greatest principal membrane curvature to orient cell wall synthesis. eLife 7:e32471 (2018).
Eun Y.J., Ho P.Y., Kim M., LaRussa S., Robert L., Renner L.D., Schmid A., Garner E.C., Amir A. Archaeal cells share common size control with bacteria despite noisier growth and division. Nature Microbiology 3(2):148-154 (2017).
Wong F., Renner L.D., Özbaykal G., Paulose J., Weibel D.B., van Teeffelen S., Amir A. Mechanical strain sensing implicated in cell shape recovery in Escherichia coli. Nature Microbiology 2:17115 (2017).
Renner L.D., Zan J., Hu L.I., Martinez M., Resto P.J., Siegel A.C., Torres C., Hall S.B., Slezak T.R., Nguyen T.H., Weibel D.B. Detection of ESKAPE bacterial pathogens at the point of care using isothermal DNA-based assays in a portable degas-actuated microfluidic diagnostic assay platform. Applied and environmental microbiology 83 (4), e02449-16 (2017).
Renner L.D., Weibel D.B. Physicochemical regulation of biofilm formation. MRS bulletin, 36 (5), 347-355 (2011)
Renner L.D., Weibel D.B. Cardiolipin microdomains localize to negatively curved regions of Escherichia coli membranes. Proceedings of the National Academy of Sciences 108 (15), 6264-6269 (2011)
